You are using an out of date browser. It may not display this or other websites correctly.
You should upgrade or use an alternative browser.
You should upgrade or use an alternative browser.
Join TeamCFC saving the World HERE!
- Thread starter Abaddon
- Start date
- Status
- Not open for further replies.
Well, you could have just a few people where each person is controlling something like 50 machines and has all 50 set up to record that one name.
For example, suppose 4 IT managers, all at different company campuses, from company X decided to start folding. The team name is the company name, but each campus folds under the same username, Campus (number), and has dozens or hundreds of machines each. That's only four users on the team as far as the stats go, but they'd be a powerhouse team.
For example, suppose 4 IT managers, all at different company campuses, from company X decided to start folding. The team name is the company name, but each campus folds under the same username, Campus (number), and has dozens or hundreds of machines each. That's only four users on the team as far as the stats go, but they'd be a powerhouse team.
VRWCAgent said:Well, you could have just a few people where each person is controlling something like 50 machines and has all 50 set up to record that one name.
For example, suppose 4 IT managers, all at different company campuses, from company X decided to start folding. The team name is the company name, but each campus folds under the same username, Campus (number), and has dozens or hundreds of machines each. That's only four users on the team as far as the stats go, but they'd be a powerhouse team.
Ah k i see what you mean
Moderator Action: Poll merged with main thread
Man, I missed the whole "breaking the top 2000" thing.  But, we're about to go over 100,000 total points, so that's cool too.
But, we're about to go over 100,000 total points, so that's cool too.  Looks like a month or so till we break the top 1000, at this rate. Good to see all the new teammates!
Looks like a month or so till we break the top 1000, at this rate. Good to see all the new teammates!
 But, we're about to go over 100,000 total points, so that's cool too.
But, we're about to go over 100,000 total points, so that's cool too.  Looks like a month or so till we break the top 1000, at this rate. Good to see all the new teammates!
Looks like a month or so till we break the top 1000, at this rate. Good to see all the new teammates!Sophie 378
Avvie by ybbor
More protein structure spam info (spoilered for the science-phobic)!
Oh, and we're on page 7 of FAHSTATS, now, not page 8:  1,909th!
1,909th!  http://fahstats.com/?col=3&offset=1501
http://fahstats.com/?col=3&offset=1501

Spoiler :
My edits etc in bold.
From http://www.steve.gb.com/science/spectroscopy.html#xray
As I attempted to explain earlier, Xray crystallography is the normal method of solving structures - biomolecules such as proteins, DNA etc are well suited to this. This explains it more concisely than the link I posted earlier - and has prettier pictures.
From http://www.steve.gb.com/science/spectroscopy.html#xray
As I attempted to explain earlier, Xray crystallography is the normal method of solving structures - biomolecules such as proteins, DNA etc are well suited to this. This explains it more concisely than the link I posted earlier - and has prettier pictures.
X-ray crystallography is the most difficult of the techniques so far described to grasp, as it is intensely mathematical. What we would really like to do with X-rays is to take pictures of atoms, like we can take pictures of bigger things with visible light. The reason we must use X-rays is that to 'see' something with light, the light must have a wavelength similar to (or smaller than) the size of the thing we are looking at. X-rays fit the bill here (visible light is far too big to see atoms), but they unfortunately cannot be focussed by any current technique. Hence we must rely on making diffraction patterns from beautifully ordered piles of molecules (crystals), rather than looking at the molecules directly. Crystallisation of the compound of interets is the primary problem in X-ray crystallography: making perfect crystals is rather fraught. Once we have a nice crystal of the compound (e.g. a protein), we illuminate it with X-rays and collect the diffracted X-rays on a photographic film, to form a diffraction pattern, like the one below.

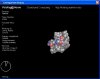
The diffraction patterns can be mathematically transformed back into electron density maps of the molecules in the crystals by Fourier transform techniques we won't go into here. A further problem of X-ray diffraction is that it relies on the size of the atoms, and hydrogen is too small to show up (Hydrogen makes up 63% of the body, so that's an important omission = but you can mostly work out where the Hs should be), hence when we 'join the dots' in our electron density map, we get something akin to a line-bond notation of the compound we have 'photographed'.

Haem electron density map: note the 'dot-to-dot' and missing hydrogens. Left, top view of unbound haem; right, side-on view on haem coordinating to two Histidines from whatever protein the haem is bound in. Carbons yellow, oxygens red, nitrogens blue, the iron atom is shown in white.
 1,909th!
1,909th!  http://fahstats.com/?col=3&offset=1501
http://fahstats.com/?col=3&offset=1501 
ironduck
Deity
- Joined
- Oct 13, 2002
- Messages
- 6,561
45 members have submitted results now! Welcome newcomers 
Sophie, the modeling from those diffraction patterns reminds me of the way engineers are getting around the wavelength limit of computer chips today (when they use optical lithography in the manufacturing) by adjusting for the diffraction patterns that appear on the mask.

Sophie, the modeling from those diffraction patterns reminds me of the way engineers are getting around the wavelength limit of computer chips today (when they use optical lithography in the manufacturing) by adjusting for the diffraction patterns that appear on the mask.
Chukchi Husky
Lone Wolf
They can be split, but it still won't make any sense afterwards.
I have another computer running, but it may take a while for it to do a work unit. It's less than half the power of mine.
I have another computer running, but it may take a while for it to do a work unit. It's less than half the power of mine.
Mathilda
Queen
I've done three WU's no problems.
Now I've got this massive one that says I'll finish it in March 2007
(providing I leave my computer on all the time)
And yes, it seems to be accurate
I don't have enough patience for one and I don't trust my computer enough to stay alive long enough to finish this.
So gurus, can I swap it for a smaller one somehow?
Now I've got this massive one that says I'll finish it in March 2007

(providing I leave my computer on all the time)
And yes, it seems to be accurate

I don't have enough patience for one and I don't trust my computer enough to stay alive long enough to finish this.
So gurus, can I swap it for a smaller one somehow?
Attachments
Yes, you can, but first I'd suggest letting it complete a couple frames of the 20000, to get a better read on it. How long has it been chewing without completing the first frame?
Mathilda
Queen
About two hours. (Give or take half an hour). I sat here last night for almost half an hour staring at it, counting down the seconds with it, and did give the impression of being accurate.
edit: forget it. You were right. It jumped straight from none to 201 and is now zooming through them at 13 seconds / frame.
edit: forget it. You were right. It jumped straight from none to 201 and is now zooming through them at 13 seconds / frame.
ironduck
Deity
- Joined
- Oct 13, 2002
- Messages
- 6,561
I still don't understand why it is that when I get a WU of a few frames each frame takes much longer than if I get one with a lot of frames.
I just finished one of 3500 frames, and each frame was around 7 seconds.
Now I'm working on one that's 400 frames and each frame is 7 minutes.
It always does this on my computer, the small ones must be really complicated and the big ones really simple?
I just finished one of 3500 frames, and each frame was around 7 seconds.
Now I'm working on one that's 400 frames and each frame is 7 minutes.
It always does this on my computer, the small ones must be really complicated and the big ones really simple?
ironduck
Deity
- Joined
- Oct 13, 2002
- Messages
- 6,561
BCLG100 said:i dont think that is the case as i completed a 3750 one and am nearly completed a 1000 one so i dont think one is more complex than the other as there is little difference in times.
I'm telling you what happens on my computer, I can't say for anyone else.
Furiey
No Longer Just Lurking
The number of points you get for a Work Unit seems to be a better measure of how long it's likely to take - they do after all try and calibrate them so that the projects are worth roughly the same number of points/time spent.
Some of the projects use higher precision calculation than others, the cores also report the results differently, some report frames, others steps. the current projects summary page will tell you what yours is worth.
Some of the projects use higher precision calculation than others, the cores also report the results differently, some report frames, others steps. the current projects summary page will tell you what yours is worth.
lost_civantares
[Insert Title]
- Joined
- Aug 30, 2005
- Messages
- 1,803
For those scared at how long a wu will take, after just finishing a WU my computer told me that the next WU would take 10987d, 21h, at 729110254 min/frame!  (hope it will be a high point WU when finished in 3985 AD!), but than changed to a much more reasonable 2 days (in accordence with windows in general it seems...).
(hope it will be a high point WU when finished in 3985 AD!), but than changed to a much more reasonable 2 days (in accordence with windows in general it seems...).
 (hope it will be a high point WU when finished in 3985 AD!), but than changed to a much more reasonable 2 days (in accordence with windows in general it seems...).
(hope it will be a high point WU when finished in 3985 AD!), but than changed to a much more reasonable 2 days (in accordence with windows in general it seems...).BCLG100 said:Ive also noticed we have moved up a lot of places very quikely in the last couple of days lets hope it continues
Yep, the only team rising faster than us nearby is Democratic Underground, apparently also a forum with just over 80,000 members, but they've got nearly double the number of folders on the team.
Abaddon
Deity
Yeah, but we have a few members with multiple computers at their bec and call 
I hope our membership levels will continue to grow, certainly with the start of a new thread we got a big increase.

I hope our membership levels will continue to grow, certainly with the start of a new thread we got a big increase.
- Status
- Not open for further replies.
Similar threads
- Replies
- 55
- Views
- 4K
- Replies
- 242
- Views
- 30K
- Replies
- 2
- Views
- 968